
The instrument can automatically set up and carry out all amplification and purification. Extracted RNA (1000 ng) with an appropriate volume of RT mix and nuclease-free water were added into the cassette, which was processed by the iR-Processor. One disposable cassette is for one sample’s library preparation all necessary reagents for amplification and purification are preloaded into the cassette. All seven chains were amplified in a single assay using a strategy which allows the incorporation of unique molecular identifiers (UMIs) during the reverse transcription (RT) step. 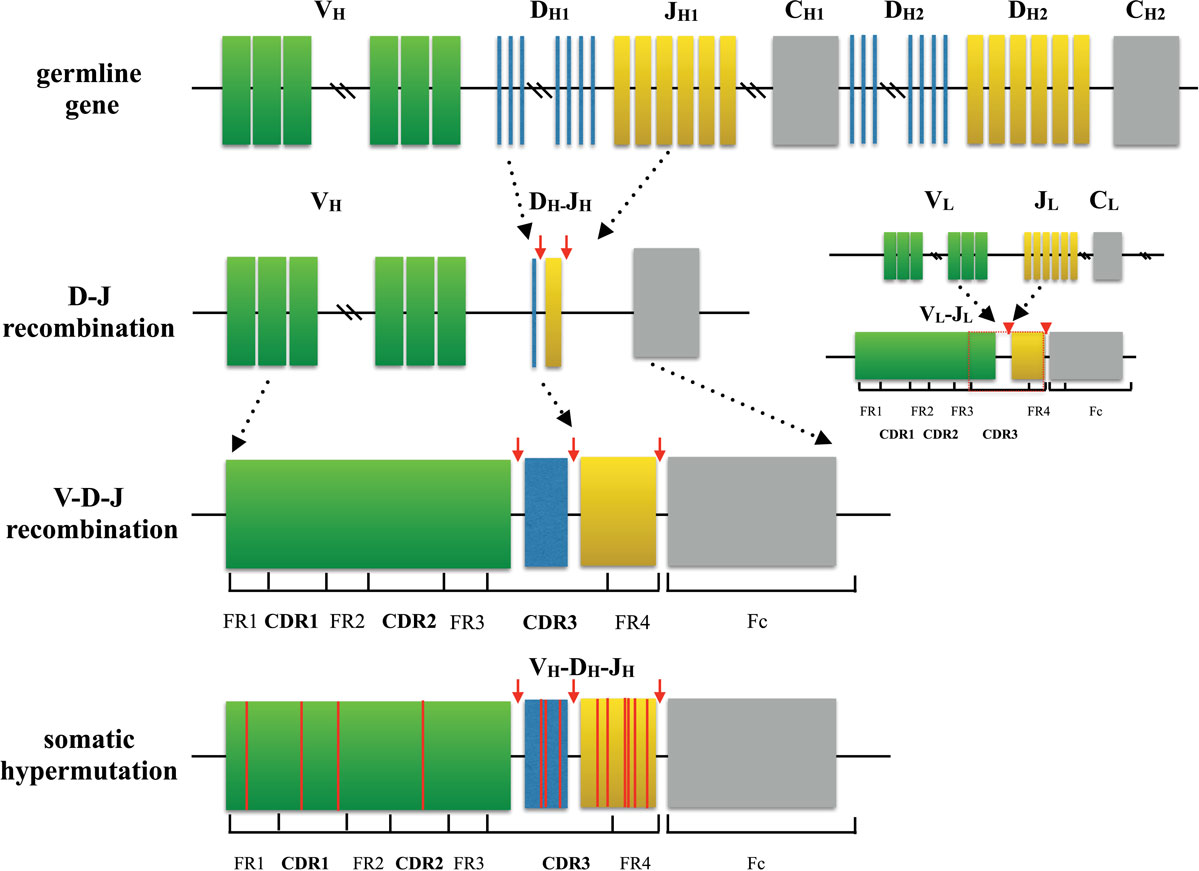
iR+7chain-HLRI-C) was used to generate NGS libraries covering all TCR and BCR chains including TCR-beta, -alpha, -delta, -gamma, and BCR-IgH, -IgK, -IgL. In this study, iR-RepSeq-plus 7-Chain Cassette (iRepertoire, catalog no. Here, we deeply investigated the peripheral blood repertoire from patients throughout their course of COVID-19 disease, demonstrating dynamic changes over the disease course. Because all seven of the immune repertoire chains, including IgH (all isotypes), IgK, IgL, TCR-Alpha, Beta chains, and TCR – Gamma, Delta chains are amplified under the same conditions in one PCR reaction with our method, the expression level of these genes can be directly compared with an inclusive and quantitative pattern ( 16). Analyzing the full repertoires could provide a better understanding of the immune response to SARS-CoV-2 and other infections. Recently, next-generation sequencing (NGS) of BCRs and TCRs have been used widely to evaluate immunity ( 17– 21). The human adaptive immune system consists of both naïve and memory cells, which express either cell surface B cell receptors (BCRs) or T cells receptors (TCRs), in aggregate termed the adaptome ( 16). The diminished T cell repertoire and progressive defects in T cell and B cell function in older patients could limit viral clearance and prolong the innate proinflammatory response ( 14, 15). Older COVID-19 patients with comorbidities are at a particularly high risk of severe pneumonia and death. Monitoring the dynamics of lymphocyte number and phenotype has been suggested as a means to predict the severity of COVID-19 ( 13). It has been observed that the counts of total T cells, CD4+ and CD8+ subtype T cells, were dramatically reduced in severe COVID-19 cases with increased expression of programmed death-ligand 1 (PD-1) and T cell immunoglobulin mucin 3 (Tim-3), indicating activation and T cell exhaustion ( 12). Lymphopenia is common in SARS-CoV-2 infected patients and was found as well as in SARS-CoV and Middle East respiratory syndrome (MERS) patients ( 4, 6, 11). Pathologic examination reveals diffuse alveolar damage, proteinaceous plugs, and a prominent myeloid infiltrate and a paucity of lymphocytes ( 7– 10).ĭespite global pandemic threats of COVID-19 disease, the host immune response against SARS-CoV-2 infection remains poorly understood. However, some COVID-19 patients rapidly develop severe pneumonia, subsequent multi-organ failure, and death ( 6). The majority of COVID-19 patients show mild or moderate symptoms and recover after proper clinical care. The pneumonia in those with this disease is characterized by bilateral ground-glass opacities identified on chest CT scans ( 4, 5). Clinical symptoms of SARS-CoV-2 include fever, cough, shortness of breath, and chest pain. The virus spreads by respiratory droplets, urine, and feces ( 4, 5). The virus caused low respiratory tract pneumonia, but it also affects multiple organs such as the kidney, liver, brain, gastrointestinal tract, and heart. The virus, severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), is a virus closely related to SARS-CoV, endemic in 2003 ( 3). 
The current outbreak of coronavirus disease (COVID-19) was first reported in Wuhan, China ( 1, 2). Profound changes in T cell homeostasis raise critical questions about the early events in COVID-19 infection and demonstrate that immune repertoire analysis is a promising method for evaluating emergent host immunity to SARS-CoV-2 viral infection, with great implications for assessing vaccination and other immunological therapies. Dominant B cell clonal expansion with decreased diversity occurred following recovery from infection. Panoramic concurrent assessment of BCR repertoires demonstrated isotype switching and a transient but dramatic early IgA expansion. Monitoring TCR repertoires could serve as an indicative biomarker to predict disease progression and recovery. Surprisingly, TCR repertoires were markedly diminished during the early onset of severe disease but recovered during the convalescent stage. We applied a novel dimer avoidance multiplexed polymerase chain reaction next-generation sequencing assay to analyze T (TCR) and B cell receptor (BCR) repertoires. Severe COVID-19 is associated with profound lymphopenia and an elevated neutrophil to lymphocyte ratio.
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
